Biology:Propionyl-CoA carboxylase
| Propionyl-CoA carboxylase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
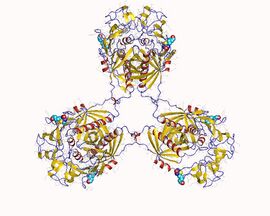
 Propionyl-CoA carboxylase hetero12mer, Methylorubrum extorquens | |||||||||
| Identifiers | |||||||||
| EC number | 6.4.1.3 | ||||||||
| CAS number | 9023-94-3 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Propionyl-CoA carboxylase (EC 6.4.1.3, PCC) catalyses the carboxylation reaction of propionyl-CoA in the mitochondrial matrix. PCC has been classified both as a ligase[1] and a lyase.[2] The enzyme is biotin-dependent. The product of the reaction is (S)-methylmalonyl CoA.
- ATP + propionyl-CoA + HCO3− <=> ADP + phosphate + (S)-methylmalonyl-CoA
(S)-Methylmalonyl-CoA cannot be directly utilized by animals. It is acted upon by a racemase, yielding (R)-methylmalonyl-CoA, which is then converted into succinyl-CoA by methylmalonyl-CoA mutase (one of the few metabolic enzymes which requires vitamin B12 as a cofactor). Succinyl-CoA, a Krebs cycle intermediate, is further metabolized into fumarate, then malate, and then oxaloacetate. Oxaloacetate may be transported into the cytosol to form phosphoenol pyruvate and other gluconeogenic intermediates. Propionyl-CoA is therefore an important precursor to glucose.
Propionyl-CoA is the end product of odd-chain fatty acid metabolism, including most methylated fatty acids. The amino acids valine, isoleucine, and methionine are also substrates for propionyl-CoA metabolism.
Structure
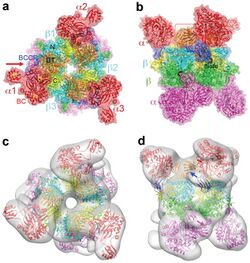
Propionyl-CoA carboxylase (PCC) is a 750 kDa alpha(6)-beta(6)-dodecamer. (Only approximately 540 kDa is native enzyme.[3] ) The alpha subunits are arranged as monomers, decorating the central beta-6 hexameric core. Said core is oriented as a short cylinder with a hole along its axis.
The alpha subunit of PCC contains the biotin carboxylase (BC) and biotin carboxyl carrier protein (BCCP) domains. A domain known as the BT domain is also located on the alpha subunit and is essential for interactions with the beta subunit. The 8-stranded anti-parallel beta barrel fold of this domain is particularly interesting. The beta subunit contains the carboxyltransferase (CT) activity.[4]
The BC and CT sites are approximately 55 Å apart, indicative of the entire BCCP domain translocating during catalysis of the carboxylation of propionyl-CoA.[5] This provides clear evidence of crucial dimeric interaction between alpha and beta subunits.
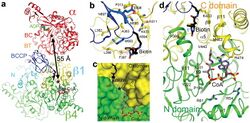
The biotin-binding pocket of PCC is hydrophobic and highly conserved. Biotin and propionyl-CoA bind perpendicular to each other in the oxyanion hole-containing active site. The native enzyme to biotin ratio has been determined to be one mole native enzyme to 4 moles biotin.[3] The N1 of biotin is thought to be the active site base.[4]
Site-directed mutagenesis at D422 shows a change in the substrate specificity of the propionyl-CoA binding site, thus indicating this residue's importance in PCC's catalytic activity.[6] In 1979, inhibition by phenylglyoxal determined that a phosphate group from either propionyl-CoA or ATP reacts with an essential arginine residue in the active site during catalysis.[7] Later (2004), it was suggested that Arginine-338 serves to orient the carboxyphosphate intermediate for optimal carboxylation of biotin.[8]
The KM values for ATP, propionyl-CoA, and bicarbonate has been determined to be 0.08 mM, 0.29 mM, and 3.0 mM, respectively. The isoelectric point falls at pH 5.5. PCC's structural integrity is conserved over the temperature range of -50 to 37 degrees Celsius and the pH range of 6.2 to 8.8. Optimum pH was shown to be between 7.2 and 8.8 without biotin bound.[3] With biotin, optimum pH is 8.0-8.5.[9]
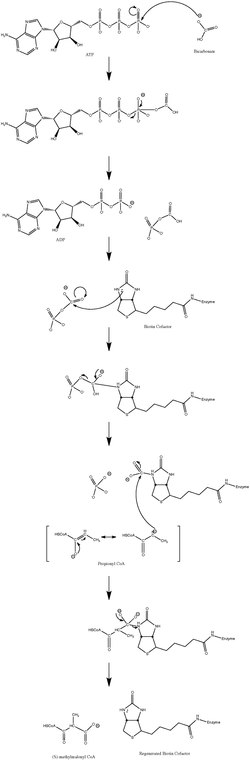
Mechanism
The normal catalytic reaction mechanism involves a carbanion intermediate and does not proceed through a concerted process.[10] Figure 3 shows a probable pathway.
The reaction has been shown to be slightly reversible at low propionyl-CoA flux.[11]
Subunit genes
Human propionyl-CoA carboxylase contains two subunits, each encoded by a separate gene:
|
| ||||||||||||||||||||||||||||||||||||||||||||||||
Pathology
A deficiency is associated with propionic acidemia.[12][13][14]
PCC activity is the most sensitive indicator of biotin status tested to date. In future pregnancy studies, the use of lymphocyte PCC activity data should prove valuable in assessment of biotin status.[15]
Intragenic complementation
When multiple copies of a polypeptide encoded by a gene form an aggregate, this protein structure is referred to as a multimer. When a multimer is formed from polypeptides produced by two different mutant alleles of a particular gene, the mixed multimer may exhibit greater functional activity than the unmixed multimers formed by each of the mutants alone. In such a case, the phenomenon is referred to as intragenic complementation.
PCC is a heteropolymer composed of α and β subunits in a α6β6 structure. Mutations in PCC, either in the α subunit (PCCα) or β subunit (PCCβ) can cause propionic acidemia in humans. When different mutant skin fibroblast cell lines defective in PCCβ were fused in pairwise combinations, the β heteromultimeric protein formed as a result often exhibited a higher level of activity than would be expected based on the activities of the parental enzymes.[16] This finding of intragenic complementation indicated that the multimeric structure of PCC allows cooperative interactions between the constituent PCCβ monomers that can generate a more functional form of the holoenzyme.
Regulation
Of Propionyl-CoA Carboxylase
a. Carbamazepine (antiepileptic drug): significantly lowers enzyme levels in the liver[17]
b. E. coli chaperonin proteins groES and groEL: essential for folding and assembly of human PCC heteromeric subunits[18]
c. Bicarbonate: negative cooperativity[8]
d. Mg2+ and MgATP2−: allosteric activation[19]
By Propionyl-CoA Carboxylase
a. 6-Deoxyerythronolide B: decrease in PCC levels lead to increased production [20]
b. Glucokinase in pancreatic beta cells: precursor of beta-PCC shown to decrease KM and increase Vmax; activation [21]
See also
References
- ↑ EC 6.4.1.3
- ↑ EC 4.1.1.41
- ↑ 3.0 3.1 3.2 "Isolation and characterization of propionyl-CoA carboxylase from normal human liver. Evidence for a protomeric tetramer of nonidentical subunits". The Journal of Biological Chemistry 255 (1): 60–65. January 1980. doi:10.1016/S0021-9258(19)86263-4. PMID 6765947.
- ↑ 4.0 4.1 "Crystal structure of the beta-subunit of acyl-CoA carboxylase: structure-based engineering of substrate specificity". Biochemistry 43 (44): 14027–14036. November 2004. doi:10.1021/bi049065v. PMID 15518551.
- ↑ 5.0 5.1 5.2 "Crystal structure of the alpha(6)beta(6) holoenzyme of propionyl-coenzyme A carboxylase". Nature 466 (7309): 1001–1005. August 2010. doi:10.1038/nature09302. PMID 20725044.
- ↑ "Crystal structures and mutational analyses of acyl-CoA carboxylase beta subunit of Streptomyces coelicolor". Biochemistry 49 (34): 7367–7376. August 2010. doi:10.1021/bi1005305. PMID 20690600.
- ↑ "Essential arginine residues in the active sites of propionyl CoA carboxylase and beta-methylcrotonyl CoA carboxylase". Enzyme 24 (5): 302–306. 1979. doi:10.1159/000458679. PMID 510274.
- ↑ 8.0 8.1 "Kinetic characterization of mutations found in propionic acidemia and methylcrotonylglycinuria: evidence for cooperativity in biotin carboxylase". The Journal of Biological Chemistry 279 (16): 15772–15778. April 2004. doi:10.1074/jbc.M311982200. PMID 14960587.
- ↑ "Human propionyl CoA carboxylase: some properties of the partially purified enzyme in fibroblasts from controls and patients with propionic acidemia". Pediatric Research 13 (6): 746–751. June 1979. doi:10.1203/00006450-197906000-00005. PMID 481943.
- ↑ "Are carboxylations involving biotin concerted or nonconcerted?". The Journal of Biological Chemistry 255 (1): 236–242. January 1980. doi:10.1016/S0021-9258(19)86289-0. PMID 7350155.
- ↑ "Assessing the reversibility of the anaplerotic reactions of the propionyl-CoA pathway in heart and liver". The Journal of Biological Chemistry 278 (37): 34959–34965. September 2003. doi:10.1074/jbc.M302013200. PMID 12824185.
- ↑ "Overview of mutations in the PCCA and PCCB genes causing propionic acidemia". Human Mutation 14 (4): 275–282. 1999. doi:10.1002/(SICI)1098-1004(199910)14:4<275::AID-HUMU1>3.0.CO;2-N. PMID 10502773.
- ↑ "Propionic acidemia: mutation update and functional and structural effects of the variant alleles". Molecular Genetics and Metabolism 83 (1–2): 28–37. 2004. doi:10.1016/j.ymgme.2004.08.001. PMID 15464417.
- ↑ "Methylmalonic and propionic aciduria". American Journal of Medical Genetics. Part C, Seminars in Medical Genetics 142C (2): 104–112. May 2006. doi:10.1002/ajmg.c.30090. PMID 16602092.
- ↑ "Lymphocyte propionyl-CoA carboxylase and its activation by biotin are sensitive indicators of marginal biotin deficiency in humans". The American Journal of Clinical Nutrition 84 (2): 384–388. August 2006. doi:10.1093/ajcn/84.1.384. PMID 16895887.
- ↑ Rodríguez-Pombo P, Pérez-Cerdá C, Pérez B, Desviat LR, Sánchez-Pulido L, Ugarte M. Towards a model to explain the intragenic complementation in the heteromultimeric protein propionyl-CoA carboxylase. Biochim Biophys Acta. 2005;1740(3):489-498. doi:10.1016/j.bbadis.2004.10.009
- ↑ "The abundance and function of biotin-dependent enzymes are reduced in rats chronically administered carbamazepine". The Journal of Nutrition 132 (11): 3405–3410. November 2002. doi:10.1093/jn/132.11.3405. PMID 12421859.
- ↑ "Chaperonin-mediated assembly of wild-type and mutant subunits of human propionyl-CoA carboxylase expressed in Escherichia coli". Human Molecular Genetics 5 (3): 331–337. March 1996. doi:10.1093/hmg/5.3.331. PMID 8852656.
- ↑ "Magnesium and magnesium adenosine triphosphate activation of human propionyl CoA carboxylase and beta-methylcrotonyl CoA carboxylase". Enzyme 28 (1): 76–81. 1982. doi:10.1159/000459088. PMID 6981505.
- ↑ "Investigating the role of native propionyl-CoA and methylmalonyl-CoA metabolism on heterologous polyketide production in Escherichia coli". Biotechnology and Bioengineering 105 (3): 567–573. February 2010. doi:10.1002/bit.22560. PMID 19806677.
- ↑ "A novel glucokinase regulator in pancreatic beta cells: precursor of propionyl-CoA carboxylase beta subunit interacts with glucokinase and augments its activity". The Journal of Biological Chemistry 276 (4): 2325–2328. January 2001. doi:10.1074/jbc.C000530200. PMID 11085976.
External links
- Propionyl-CoA+Carboxylase at the US National Library of Medicine Medical Subject Headings (MeSH)
 |